pycirclize带你轻松绘制基因组图
pycirclize带你轻松绘制基因组图
R语言数据分析指南
发布于 2023-08-18 13:36:07
发布于 2023-08-18 13:36:07
欢迎关注R语言数据分析指南
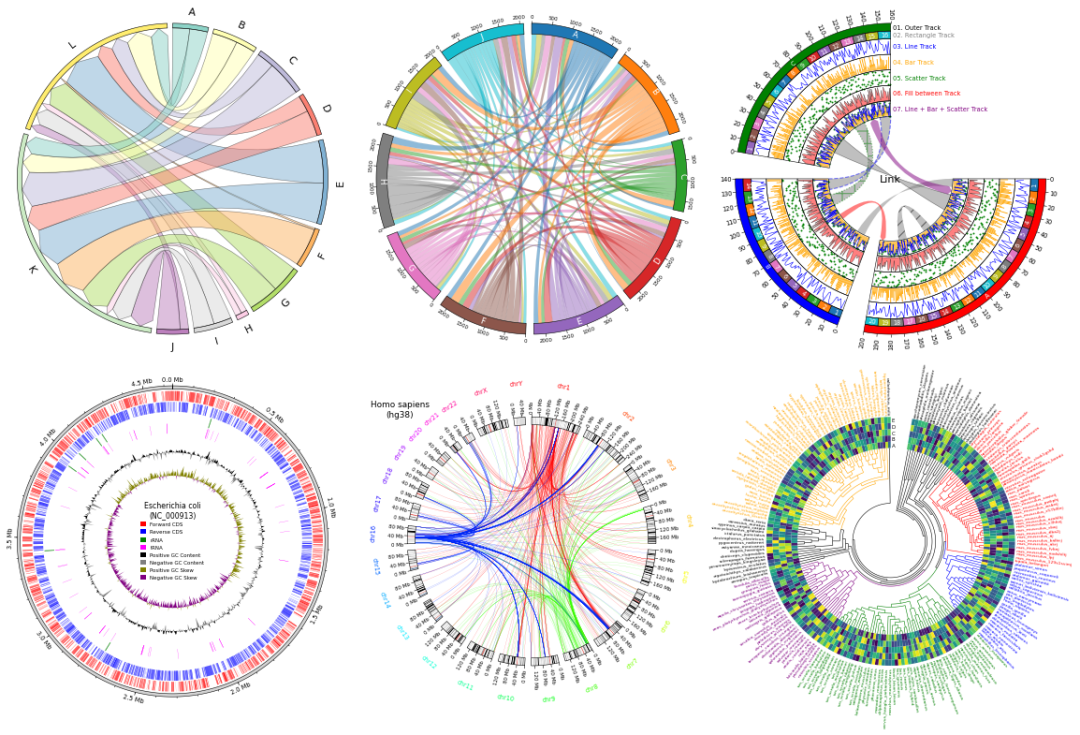
❝本节来介绍一款基于matplotlib实现的圆形可视化python包「pyCirclize」,作者结合了「circlize」&「circos」的特点功能更加的强大,结果展示也是非常的丰富。感觉非常有实用价值,小编在此展示部分内容,更多详细内容请参考作者官方文档。 ❞
官方文档
❝https://github.com/moshi4/pyCirclize https://moshi4.github.io/pyCirclize/getting_started/ ❞
安装Python包
pip install pycirclize
conda install -c conda-forge pycirclize
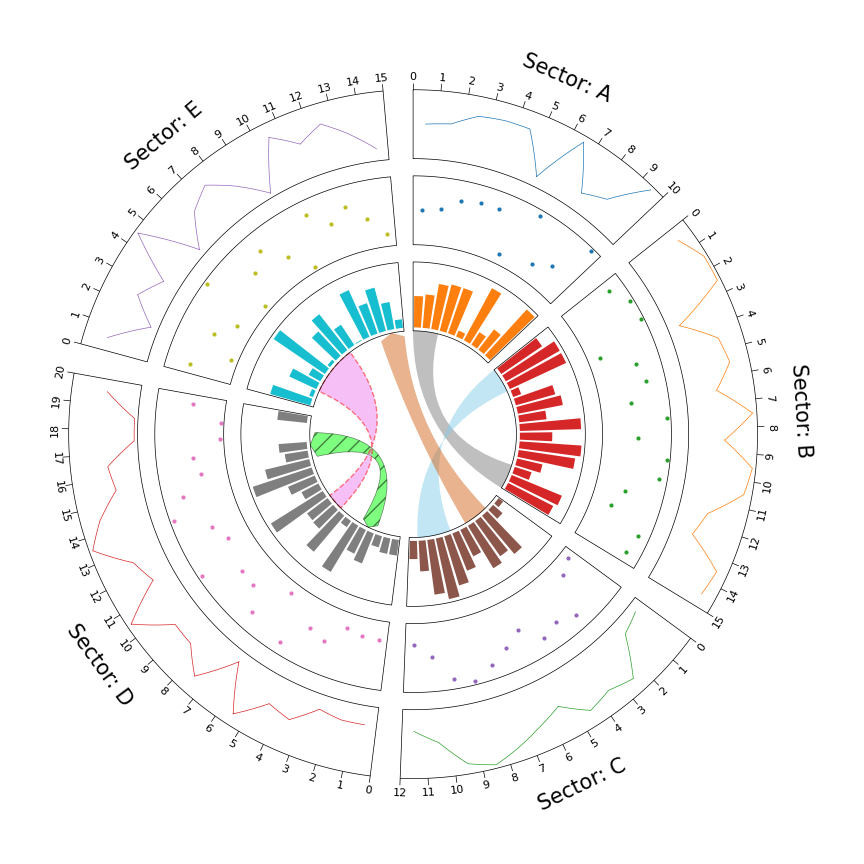
案例1 绘制和弦图
from pycirclize import Circos
import numpy as np
np.random.seed(0)
sectors = {"A": 10, "B": 15, "C": 12, "D": 20, "E": 15}
circos = Circos(sectors, space=5)
for sector in circos.sectors:
# Plot sector name
sector.text(f"Sector: {sector.name}", r=110, size=15)
# Create x positions & random y values
x = np.arange(sector.start, sector.end) + 0.5
y = np.random.randint(0, 100, len(x))
# Plot lines
track1 = sector.add_track((80, 100), r_pad_ratio=0.1)
track1.xticks_by_interval(interval=1)
track1.axis()
track1.line(x, y)
# Plot points
track2 = sector.add_track((55, 75), r_pad_ratio=0.1)
track2.axis()
track2.scatter(x, y)
# Plot bars
track3 = sector.add_track((30, 50), r_pad_ratio=0.1)
track3.axis()
track3.bar(x, y)
circos.link(("A", 0, 3), ("B", 15, 12))
circos.link(("B", 0, 3), ("C", 7, 11), color="skyblue")
circos.link(("C", 2, 5), ("E", 15, 12), color="chocolate", direction=1)
circos.link(("D", 3, 5), ("D", 18, 15), color="lime", ec="black", lw=0.5, hatch="//", direction=2)
circos.link(("D", 8, 10), ("E", 2, 8), color="violet", ec="red", lw=1.0, ls="dashed")
circos.savefig("example01.png")
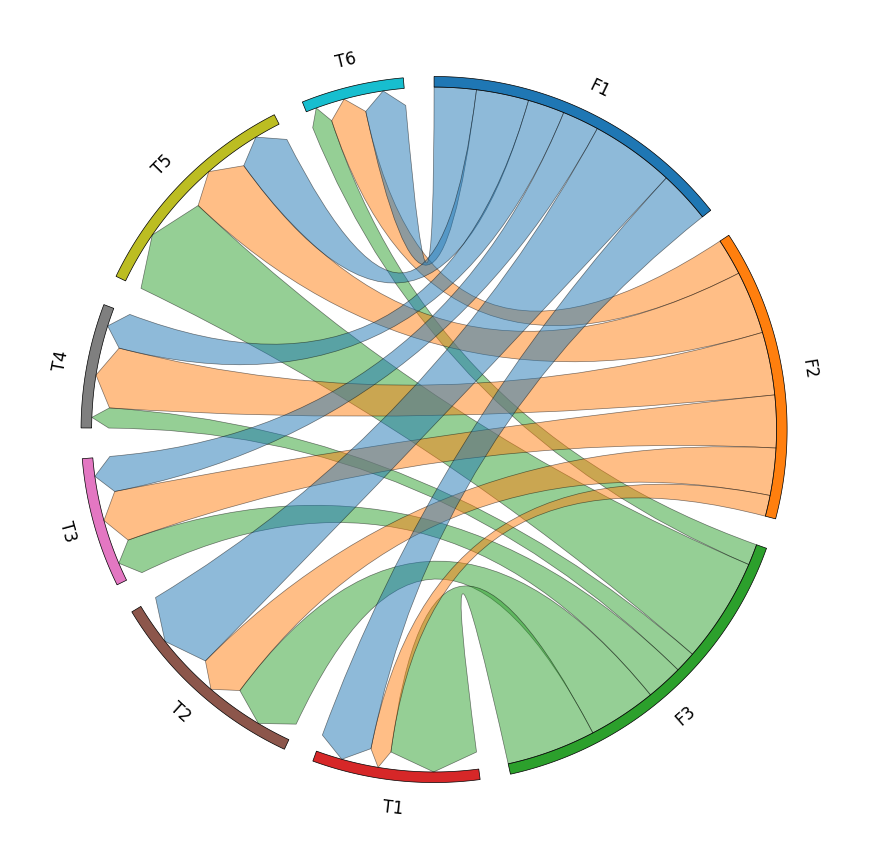
案例2 基础和弦图
from pycirclize import Circos
import pandas as pd
# Create matrix dataframe (3 x 6)
row_names = ["F1", "F2", "F3"]
col_names = ["T1", "T2", "T3", "T4", "T5", "T6"]
matrix_data = [
[10, 16, 7, 7, 10, 8],
[4, 9, 10, 12, 12, 7],
[17, 13, 7, 4, 20, 4],
]
matrix_df = pd.DataFrame(matrix_data, index=row_names, columns=col_names)
# Initialize Circos from matrix for plotting Chord Diagram
circos = Circos.initialize_from_matrix(
matrix_df,
space=5,
cmap="tab10",
label_kws=dict(size=12),
link_kws=dict(ec="black", lw=0.5, direction=1),
)
circos.savefig("example02.png")
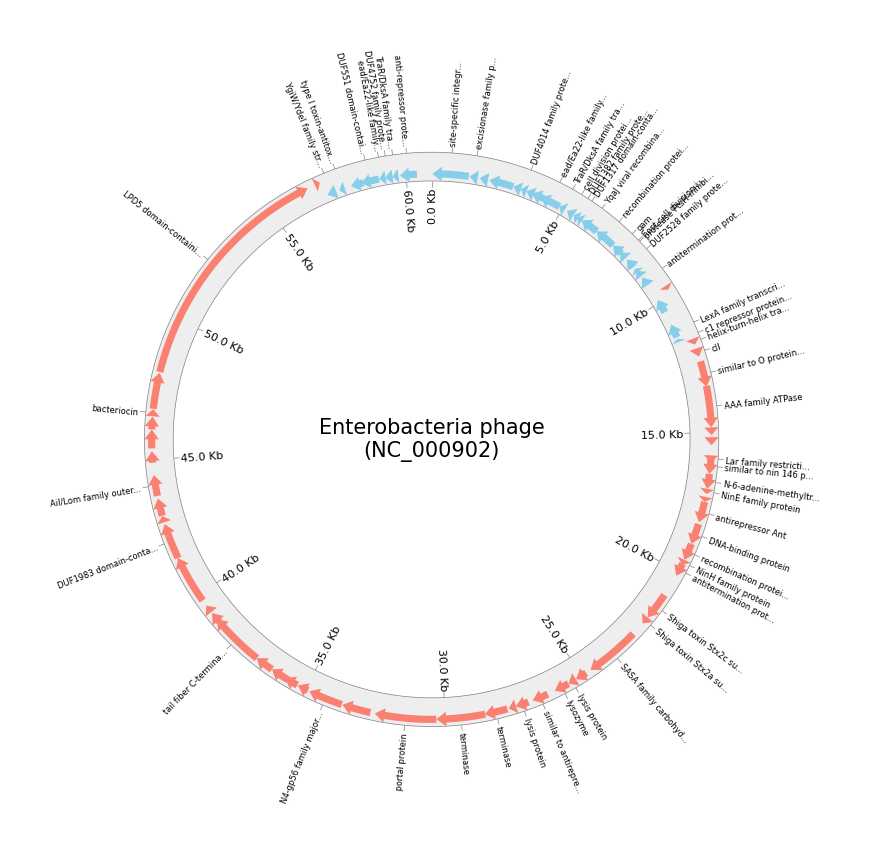
案例3 绘制基因组图
from pycirclize import Circos
from pycirclize.parser import Gff
from pycirclize.utils import load_prokaryote_example_file
# Load GFF file
gff_file = load_prokaryote_example_file("enterobacteria_phage.gff")
gff = Gff(gff_file)
circos = Circos(sectors={gff.name: gff.range_size})
circos.text("Enterobacteria phage\n(NC_000902)", size=15)
sector = circos.sectors[0]
cds_track = sector.add_track((90, 100))
cds_track.axis(fc="#EEEEEE", ec="none")
# Plot forward CDS
cds_track.genomic_features(
gff.extract_features("CDS", target_strand=1),
plotstyle="arrow",
r_lim=(95, 100),
fc="salmon",
)
# Plot reverse CDS
cds_track.genomic_features(
gff.extract_features("CDS", target_strand=-1),
plotstyle="arrow",
r_lim=(90, 95),
fc="skyblue",
)
# Extract CDS product labels
pos_list, labels = [], []
for feat in gff.extract_features("CDS"):
start, end = int(str(feat.location.end)), int(str(feat.location.start))
pos = (start + end) / 2
label = feat.qualifiers.get("product", [""])[0]
if label == "" or label.startswith("hypothetical"):
continue
if len(label) > 20:
label = label[:20] + "..."
pos_list.append(pos)
labels.append(label)
# Plot CDS product labels on outer position
cds_track.xticks(
pos_list,
labels,
label_orientation="vertical",
show_bottom_line=True,
label_size=6,
line_kws=dict(ec="grey"),
)
# Plot xticks & intervals on inner position
cds_track.xticks_by_interval(
interval=5000,
outer=False,
show_bottom_line=True,
label_formatter=lambda v: f"{v/ 1000:.1f} Kb",
label_orientation="vertical",
line_kws=dict(ec="grey"),
)
fig = circos.plotfig()
绘制人基因组图
from pycirclize import Circos
from pycirclize.utils import load_eukaryote_example_dataset
# Load hg38 dataset (https://github.com/moshi4/pycirclize-data/tree/main/eukaryote/hg38)
chr_bed_file, cytoband_file, _ = load_eukaryote_example_dataset("hg38")
# Initialize Circos from BED chromosomes
circos = Circos.initialize_from_bed(chr_bed_file, space=3)
circos.text("Homo sapiens (hg38)", size=15)
# Add cytoband tracks from cytoband file
circos.add_cytoband_tracks((95, 100), cytoband_file)
# Plot chromosome name
for sector in circos.sectors:
sector.text(sector.name, size=10)
fig = circos.plotfig()
from pycirclize import Circos
from pycirclize.utils import ColorCycler, load_eukaryote_example_dataset
# Load hg38 dataset (https://github.com/moshi4/pycirclize-data/tree/main/eukaryote/hg38)
chr_bed_file, cytoband_file, chr_links = load_eukaryote_example_dataset("hg38")
# Initialize Circos from BED chromosomes
circos = Circos.initialize_from_bed(chr_bed_file, space=3)
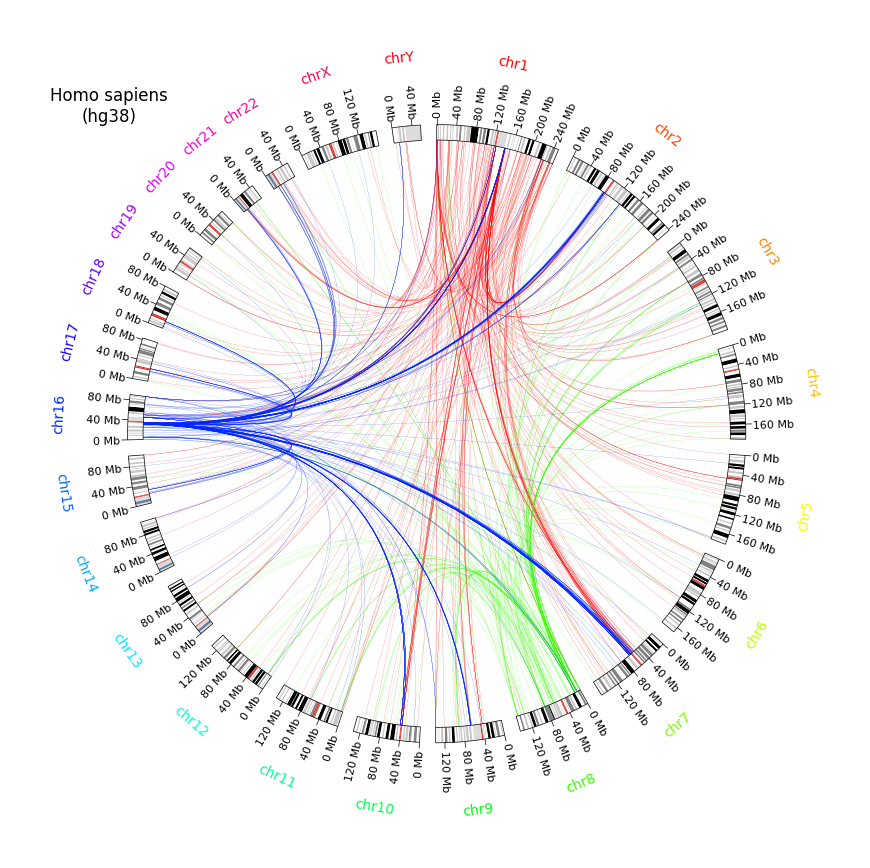
circos.text("Homo sapiens\n(hg38)", deg=315, r=150, size=12)
# Add cytoband tracks from cytoband file
circos.add_cytoband_tracks((95, 100), cytoband_file)
# Create chromosome color mapping
ColorCycler.set_cmap("hsv")
chr_names = [s.name for s in circos.sectors]
colors = ColorCycler.get_color_list(len(chr_names))
chr_name2color = {name: color for name, color in zip(chr_names, colors)}
# Plot chromosome name & xticks
for sector in circos.sectors:
sector.text(sector.name, r=120, size=10, color=chr_name2color[sector.name])
sector.get_track("cytoband").xticks_by_interval(
40000000,
label_size=8,
label_orientation="vertical",
label_formatter=lambda v: f"{v / 1000000:.0f} Mb",
)
# Plot chromosome link
for link in chr_links:
region1 = (link.query_chr, link.query_start, link.query_end)
region2 = (link.ref_chr, link.ref_start, link.ref_end)
color = chr_name2color[link.query_chr]
if link.query_chr in ("chr1", "chr8", "chr16") and link.query_chr != link.ref_chr:
circos.link(region1, region2, color=color)
fig = circos.plotfig()
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2023-06-17,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

