空转联合单细胞分析(八):Visium HD多样本整合分析(基于seurat)
空转联合单细胞分析(八):Visium HD多样本整合分析(基于seurat)

KS科研分享与服务-TS的美梦
发布于 2025-12-18 14:05:31
发布于 2025-12-18 14:05:31
本内容前期准备参考:
空转上游你可以不做,但不能不会呀!详细演示Visium HD上游公共数据库分析及图像对齐矫正
空转联合单细胞分析(五):10X Visium HD上手就是复杂情况?多样本拼片如何进行拆分?
空转联合单细胞分析(六):10X Visium HD seurat V5分析教程
空转联合单细胞分析(七):10X Visium HD基于空间信息的BANKSY聚类
之前对于HD的演示,虽然一张捕获区域有多个sample,但是我们是按照正常的流程演示,当作一张片子进行的分析。有小伙伴需求多样本的整合分析,这里演示一下,流程是类似的。
1、数据下载
关于visium HD数据的seurat读取处理以及其他情况的处理,已经有一个详细的教程了。之前的示例数据多个组在一张slice,我们是按照单个样本演示的。这里演示一下visium HD多样本整合分析,流程与普通visium是类似的。选取的数据是发表在Oliveira, M.F.d., Romero, J.P., Chung, M. et al. High-definition spatial transcriptomic profiling of immune cell populations in colorectal cancer. Nat Genet 57, 1512–1523 (2025). https://doi.org/10.1038/s41588-025-02193-3上的文章。这是一组很经典的示例数据,具体的数据可以在GEO下载,也可以在10x官网下载完整的结果或者自行下载FATSQ数据跑上游。样本中来自人类结直肠癌及其邻近正常黏膜组织的空间转录组,原本有5例样本,这里选择1例直肠癌、1例正常对照进行演示。数据下载链接:https://www.10xgenomics.com/platforms/visium/product-family/dataset-human-crc:
data download:
wget https://cf.10xgenomics.com/samples/spatial-exp/3.0.0/Visium_HD_Human_Colon_Normal_P5/Visium_HD_Human_Colon_Normal_P5_binned_outputs.tar.gz
wget https://cf.10xgenomics.com/samples/spatial-exp/3.0.0/Visium_HD_Human_Colon_Cancer_P5/Visium_HD_Human_Colon_Cancer_P5_binned_outputs.tar.gz
tar -xvzf Visium_HD_Human_Colon_Normal_P5
tar -xvzf Visium_HD_Human_Colon_Cancer_P5setwd("~/data_analysis/10X_space/CRC_visiumHD/")
library(Seurat)
library(ggplot2)
library(patchwork)
library(dplyr)2、读取数据及基本处理
我们选择8um分辨率的数据:
CRC <- Load10X_Spatial(data.dir = './CRC/binned_outputs/square_008um/',slice = 'CRC')
CRC$group <- 'CRC'#添加分组
CRC## An object of class Seurat
## 18085 features across 541968 samples within 1 assay
## Active assay: Spatial (18085 features, 0 variable features)
## 1 layer present: counts
## 1 spatial field of view present: CRCNAT <- Load10X_Spatial(data.dir = './NAT/binned_outputs/square_008um/',slice ='NAT')
NAT$group <- 'NAT' #添加分组
NAT## An object of class Seurat
## 18085 features across 435773 samples within 1 assay
## Active assay: Spatial (18085 features, 0 variable features)
## 1 layer present: counts
## 1 spatial field of view present: NAT将数据合并为一个list,用于质控:
obj.list <- list(CRC,NAT)
for (i in 1:length(obj.list)) {
obj.list[[i]][["percent.mt"]] <- PercentageFeatureSet(obj.list[[i]], pattern = "^MT-")#计算bins线粒体基因比例
}object_meta1 = obj.list[[1]]@meta.data
# Create a plot for nUMI
dist_counts_before1 <- object_meta1 %>%
ggplot(aes(x=nCount_Spatial)) +
geom_density(alpha = 0.2) +
scale_x_log10() +
theme_classic() +
ylab("Cell density") +
xlab("Number of UMIs per bin") +
ggtitle('Pre-QC UMIs/Bin') +
theme(plot.title = element_text(hjust = 0.5))
# Create a plot for nGene
dist_features_before1 <- object_meta1 %>%
ggplot(aes(x=nFeature_Spatial)) +
geom_density(alpha = 0.2) +
scale_x_log10() +
theme_classic() +
ylab("Cell density") +
xlab("Number of genes per bin") +
ggtitle('Pre-QC Genes/Bin') +
theme(plot.title = element_text(hjust = 0.5))
# Create a plot for percent.mt
dist_mt_before1 <- object_meta1 %>%
ggplot(aes(x=percent.mt)) +
geom_density(alpha = 0.2) +
scale_x_log10() +
theme_classic() +
ylab("Cell density") +
xlab("Number of genes per bin") +
ggtitle('Pre-QC pt.mt/Bin') +
theme(plot.title = element_text(hjust = 0.5))
dist_counts_before1 | dist_features_before1 | dist_mt_before1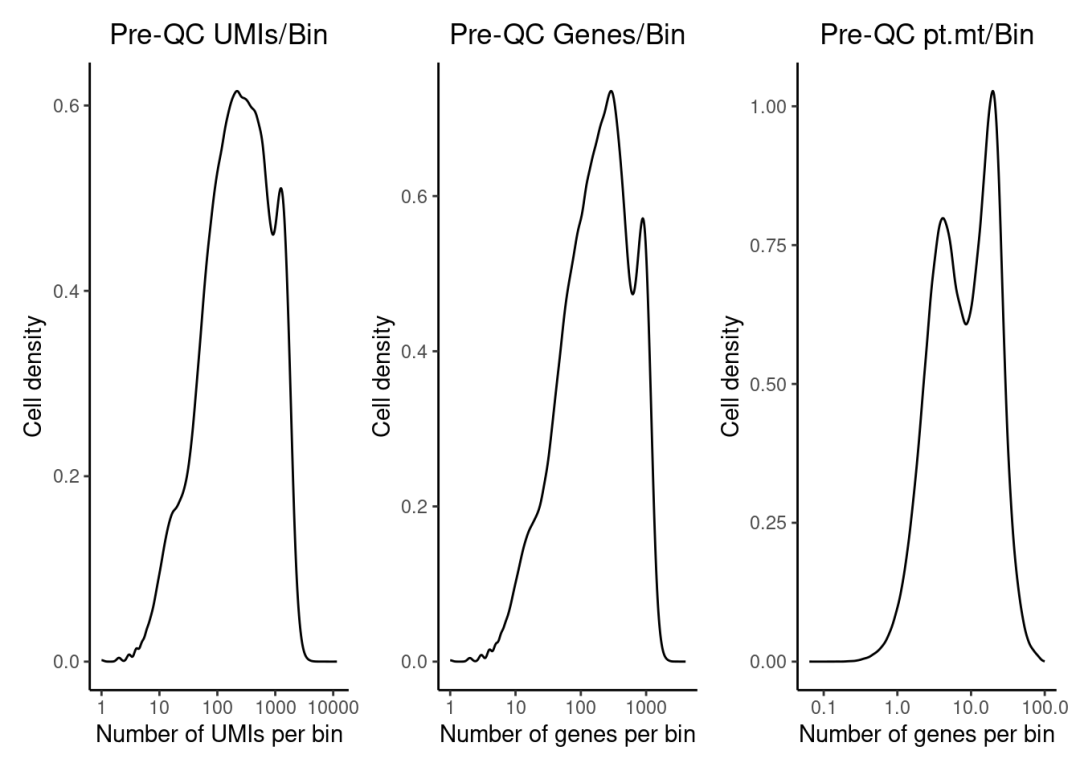
数据质控,这里我们过滤掉线粒体基因比例大于40%,UMIs和Features小于100的bins【质控阈值请根据自己实际数据调整,这不是金标准】。
for (i in 1:length(obj.list)) {
obj.list[[i]] <- subset(obj.list[[i]], subset = nCount_Spatial > 100 & nFeature_Spatial > 100 & percent.mt < 40)
}合并:
CRC_merge <- merge(obj.list[[1]],obj.list[[2]],add.cell.ids = c("CRC", "NAT"))
CRC_merge3、sketch聚类
标准化:
CRC_merge <- NormalizeData(CRC_merge, assay = 'Spatial')CRC_merge <- FindVariableFeatures(CRC_merge)CRC_merge <- SketchData(
object = CRC_merge,
assay = 'Spatial',
ncells = ncol(CRC_merge) * 0.15,
method = "LeverageScore",
sketched.assay = "sketch"
)
CRC_merge## An object of class Seurat
## 36170 features across 616313 samples within 2 assays
## Active assay: sketch (18085 features, 2000 variable features)
## 4 layers present: counts.1, counts.2, data.1, data.2
## 1 other assay present: Spatial
## 2 spatial fields of view present: CRC NAT在sketch assay对采样的细胞进行标准化降维流程以及样本整合:
DefaultAssay(CRC_merge) <- "sketch"
CRC_merge <- FindVariableFeatures(CRC_merge, verbose = FALSE)
CRC_merge <- ScaleData(CRC_merge, verbose = FALSE)
CRC_merge <- RunPCA(CRC_merge, assay = "sketch", reduction.name = "pca.sketch", verbose = FALSE)processed with batch correction using Harmony【因为visium HD数据很大,所以整合是比较费时间的,如果分析需求没有对不同样本进行对比分析的,其实可以单独分开进行分析,没必要整合。或者在下采样的时候选择的细胞数少一点】:
CRC_merge <- IntegrateLayers(object = CRC_merge,
method = HarmonyIntegration,
orig.reduction = "pca.sketch",
new.reduction = "integrated.Harmony",
verbose = FALSE)降维聚类:
CRC_merge <- FindNeighbors(CRC_merge,
assay = "sketch",
reduction = "integrated.Harmony", dims = 1:25)
CRC_merge <- FindClusters(CRC_merge,
cluster.name = "seurat_cluster.sketched",
resolution = 0.5)CRC_merge <- RunUMAP(CRC_merge, reduction = "integrated.Harmony",
reduction.name = "umap.sketch", return.model = T,
dims = 1:25)# Plot UMAP
DimPlot(CRC_merge, reduction = "umap.sketch", label = T, cols = 'polychrome') +
ggtitle("Sketched clustering") +
theme(legend.position = "none")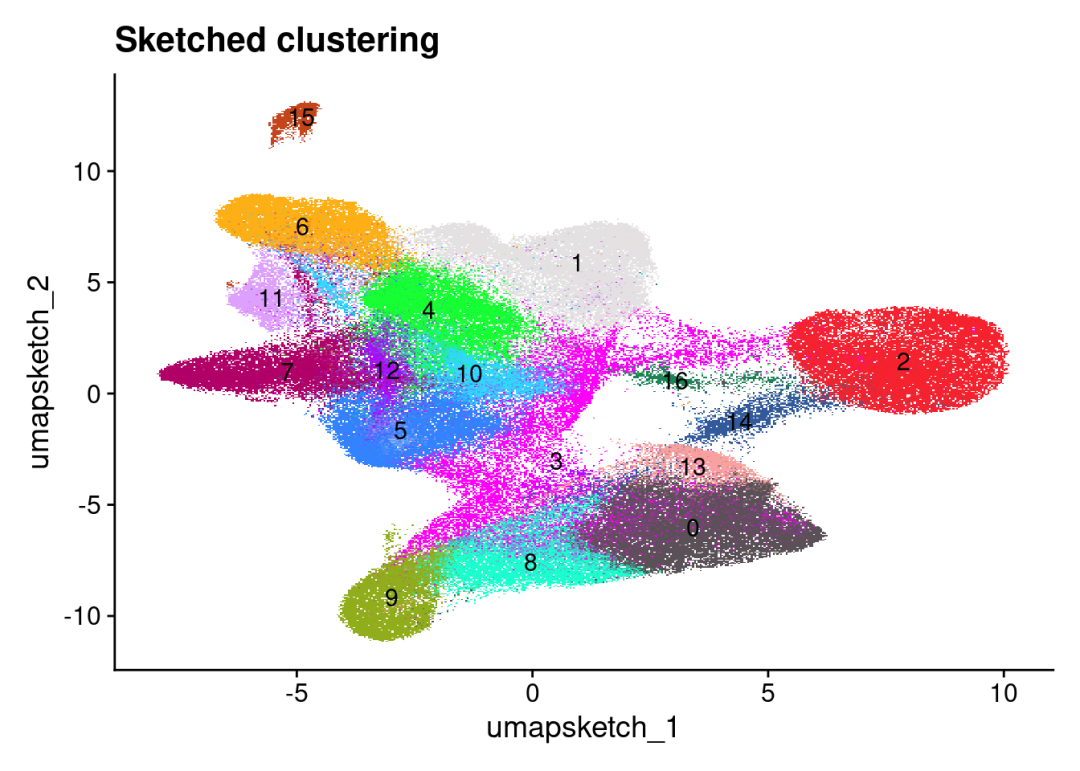
# Plot UMAP
DimPlot(CRC_merge, reduction = "umap.sketch", group.by = 'group')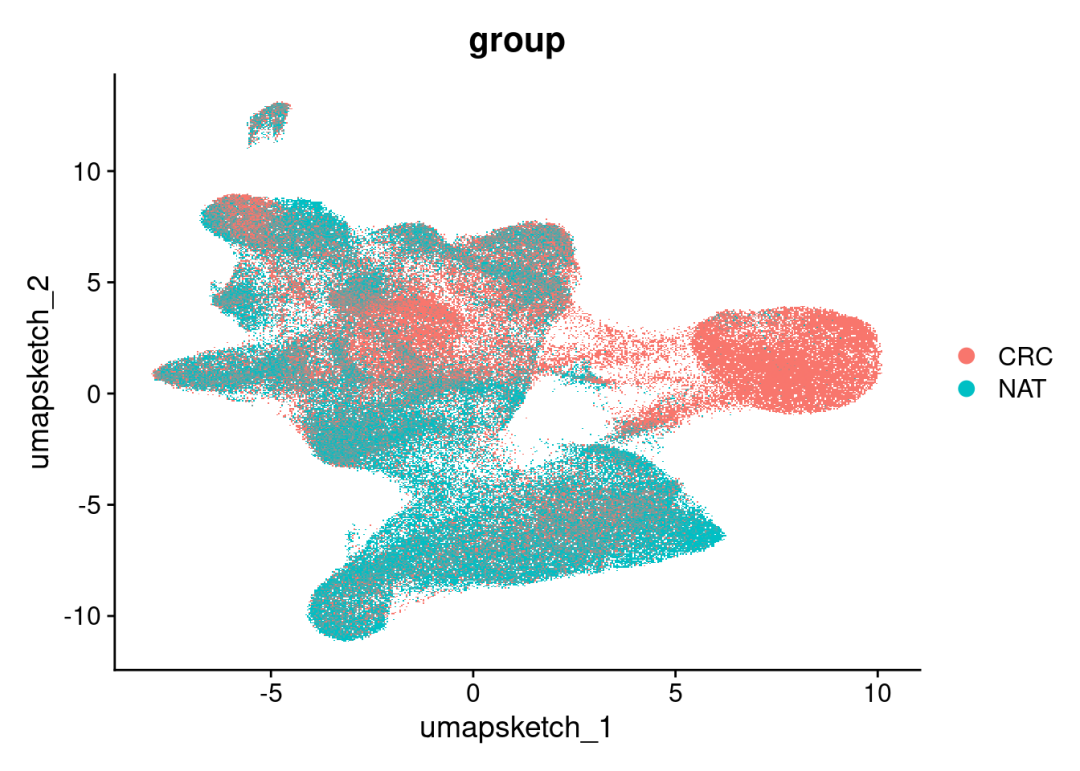
4、将sketch聚类标签投射回完整数据集
将“sketch”阶段得到的结果投射回所有 bins。这两步骤比较耗费时间!
#返回的整合后的full reduction为integrated.Harmony.full
CRC_merge <- ProjectIntegration(object = CRC_merge, #合并的seurat obj
sketched.assay = "sketch", #sketch assay
assay = "Spatial",#原始数据assay
reduction="integrated.Harmony")#使用sketch,经过批次校正的嵌入的降维结果名称。options(future.globals.maxSize = 10 * 1024^3)
CRC_merge <- ProjectData(
object = CRC_merge,
assay = "Spatial",#原来的完整数据集assay
full.reduction = "integrated.Harmony.full",#投射后所有bins的reduction名称
sketched.assay = "sketch",#sketch下采样数据assay
sketched.reduction = "integrated.Harmony.full",#投射需要使用的sketched.reduction
umap.model = "umap.sketch",#sketch上训练好的UMAP模型的名称
dims = 1:25,
refdata = list(seurat_cluster.projected = "seurat_cluster.sketched")
)## Finding sketch neighbors## Finding sketch weight matrix## Transfering refdata from sketch## Projection to sketch umap## Running UMAP projection## 12:01:50 Read 616313 rows## 12:01:50 Processing block 1 of 1## 12:01:50 Commencing smooth kNN distance calibration using 1 thread with target n_neighbors = 30
## 12:02:03 Initializing by weighted average of neighbor coordinates using 1 thread
## 12:02:08 Commencing optimization for 67 epochs, with 18489390 positive edges
## 12:04:22 Finished## Warning: Keys should be one or more alphanumeric characters followed by an
## underscore, setting key from full.umap.sketch to fullumapsketch_可视化最终效果:
DefaultAssay(CRC_merge) <- "Spatial"
Idents(CRC_merge) <- "seurat_cluster.projected"
# Plot the UMAP
DimPlot(CRC_merge, reduction = "full.umap.sketch", label = T, raster = F, cols = 'polychrome') +
ggtitle("Projected clustering") +
theme(legend.position = "none")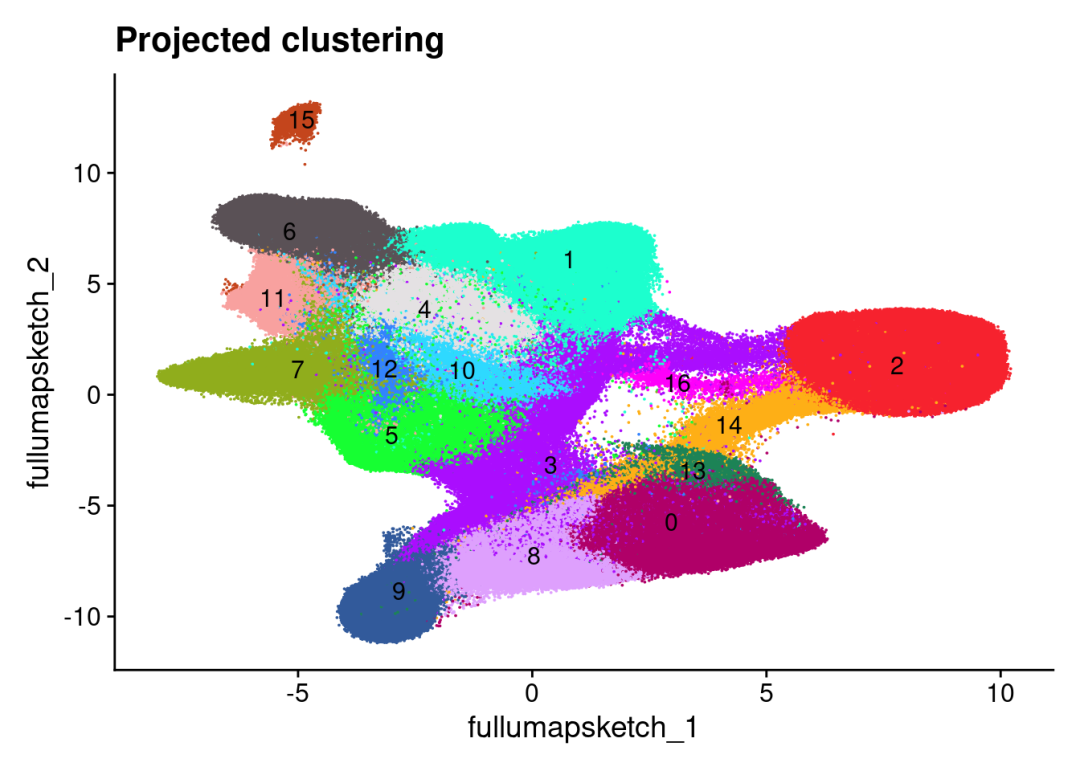
可视化组织:
color_pal=c( "#E41A1C","#377EB8","#4DAF4A", "#984EA3","#FF7F00","#FFFF33","#A65628",
"#F781BF","#999999","#8DD3C7","#FFFFB3","#BEBADA","#FB8072","#80B1D3",
"#FDB462", "#B3DE69","#FCCDE5","#CCEBC5","#D95F02","#003366","#1B9E77",
"#7570B3","#E7298A","#66A61E","#E6AB02","#FFC0CB")
SpatialDimPlot(CRC_merge,
group.by = 'seurat_cluster.projected',
pt.size.factor = 8,
images = 'CRC') +
scale_fill_manual(values = color_pal)+
guides(fill=guide_legend(ncol=2))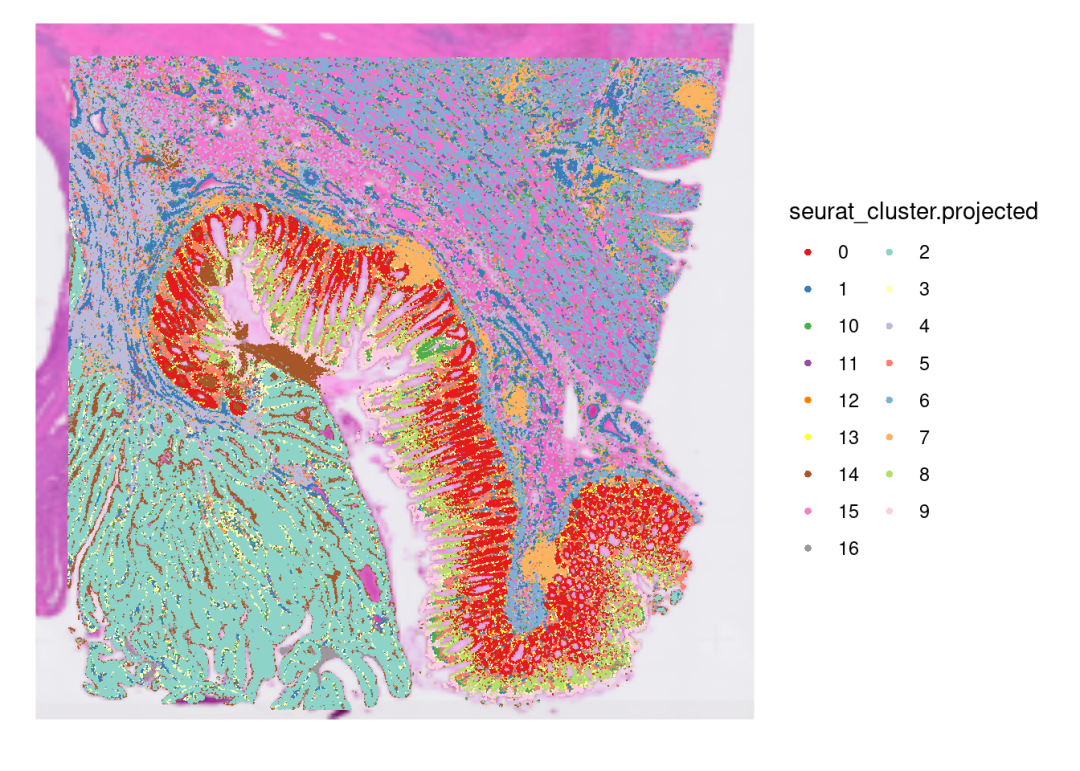
color_pal=c( "#E41A1C","#377EB8","#4DAF4A", "#984EA3","#FF7F00","#FFFF33","#A65628",
"#F781BF","#999999","#8DD3C7","#FFFFB3","#BEBADA","#FB8072","#80B1D3",
"#FDB462", "#B3DE69","#FCCDE5","#CCEBC5","#D95F02","#003366","#1B9E77",
"#7570B3","#E7298A","#66A61E","#E6AB02","#FFC0CB")
SpatialDimPlot(CRC_merge,
group.by = 'seurat_cluster.projected',
pt.size.factor = 8,
images = 'NAT') +
scale_fill_manual(values = color_pal)+
guides(fill=guide_legend(ncol=2))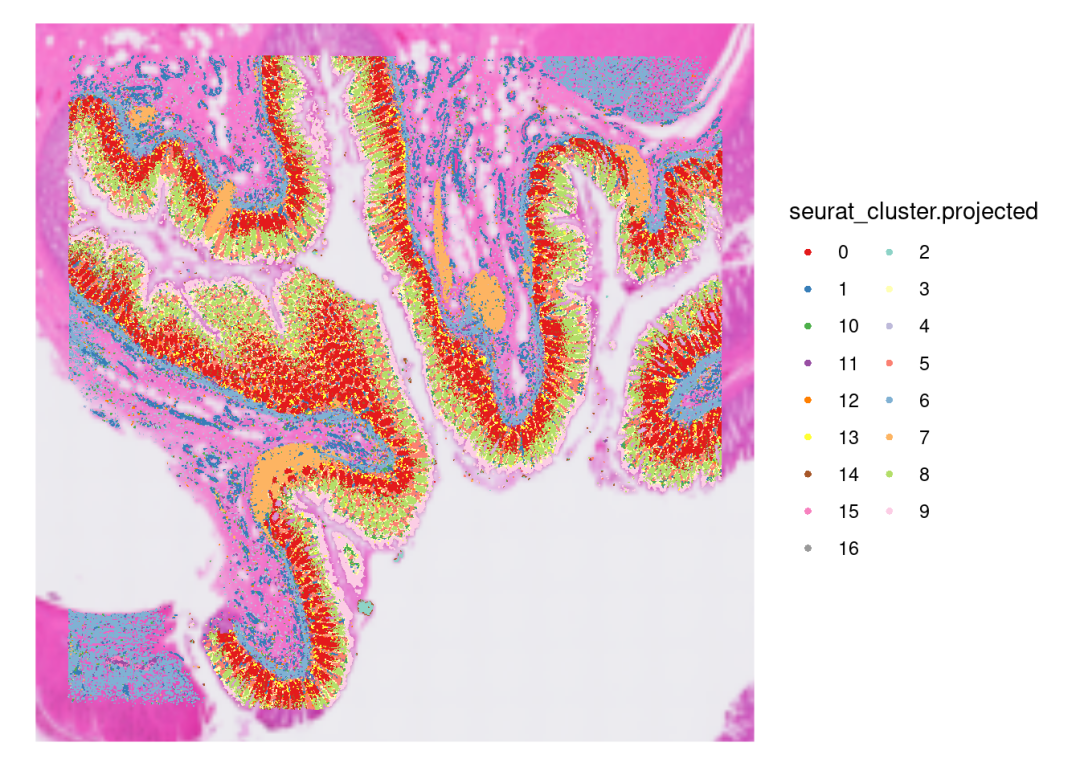
对比原文,其实整个降维后的cluster分布是有一定的特征的。原文描述未见对数据的质控,我们进行了过滤之后,可以看到有部分bins在组织上直接缺失了,所以实际处理中,对于数据的质控还是需要结数据进行设计!
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2025-12-16,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

