课前准备--基因组(WGS/WES)联合空转获取突变信息
原创课前准备--基因组(WGS/WES)联合空转获取突变信息
原创
追风少年i
发布于 2025-09-07 10:48:50
发布于 2025-09-07 10:48:50
作者,Evil Genius
生化小课马上开始了,感兴趣的可以参加一下,为自己储备一些分析技能,为单细胞空间多组学分析做准备,链接在生化小课---基因组与单细胞空间多组学
虽然本次课程主要强调的是基因组 + 单细胞空间多组学,但是空转不仅仅可以辅助基因组,还有微生物组。
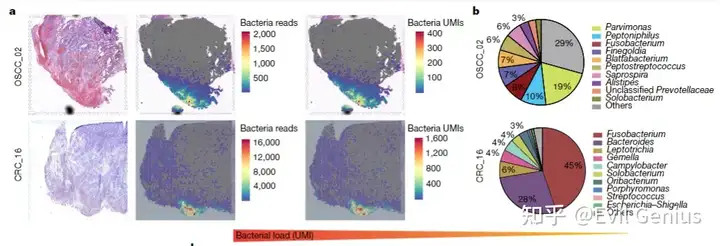
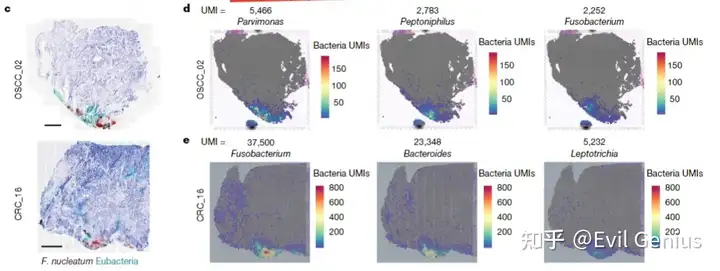
所以大家在单细胞程度上多扩展一步,目前会收到很大的效果,但是再过几年,基本上窗口期也就过了。
今天我们需要实现的目标,空间转录组检测突变
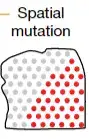
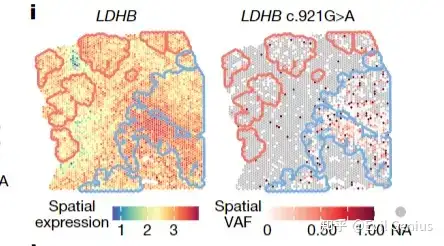
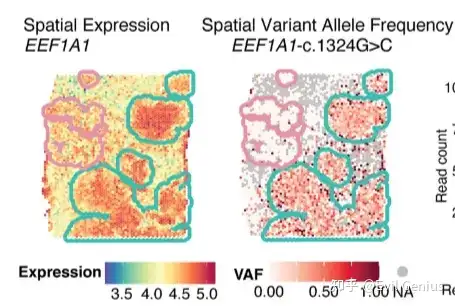
实现上面的分析严格上也需要WGS/WES + 空转两个组学的数据。
WES/WGS的分析
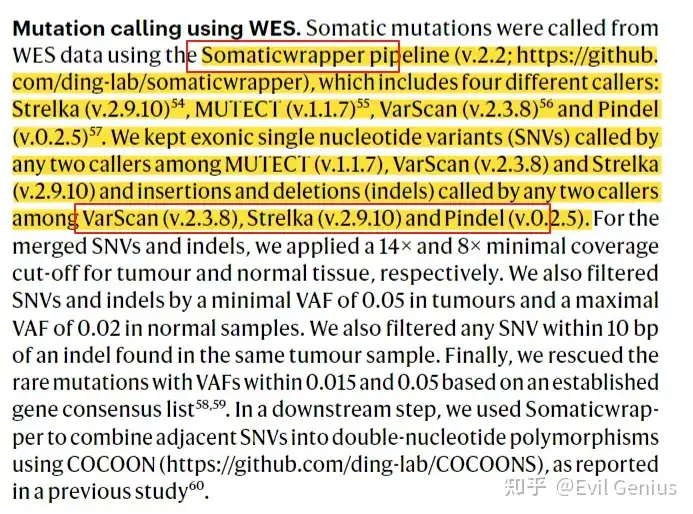

全流程自动化的基因组脚本call SNV + ANNOVAR + MSI + Fusion + CNV + TMB ,课上我会发给大家。
第二步:Mutation mapping to snRNA-seq and ST data


我们来实现一下,比较简单,脚本在https://github.com/ding-lab/10Xmapping。
#step 1: extracting reads contains the reference allele and variant allele of a somatic variant perl 10Xmapping.pl --mapq --bam --maf --out ##mapq: the mapping quality; Default 0 ##bam: the scRNA-seq bam path ##maf: the maf file containing a list of somatic variants(WES的分析结果) ##out: the output file #step 2: get the corresponding table between ref allele, var allele and 10X barcode perl parse_scrna_bc.pl $fin $fout ##fin: the input file from step 1's output ##fout: the output file #step 3: output number of reads supporting reference and variant alleles and VAF perl rc.pl $fin $fout ##fin: the input file from step 1's output ##fout: the output file
同样,会获得每个barocde对应的突变信息。
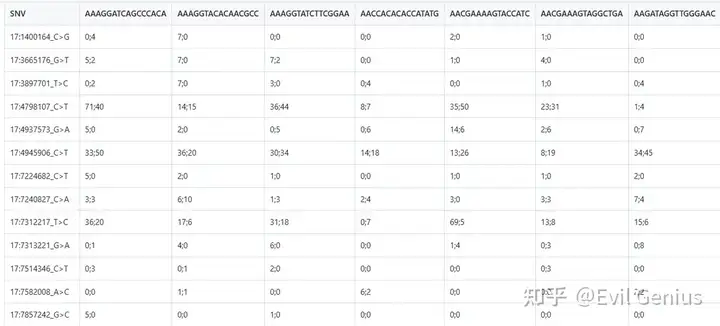
第三步可视化,这个就简单了,画一画即可
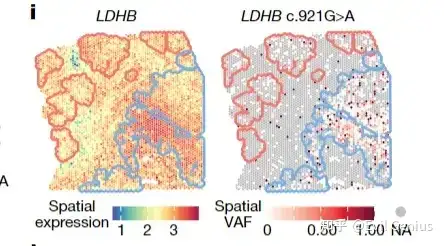
生活很好,有你更好
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
原创声明:本文系作者授权腾讯云开发者社区发表,未经许可,不得转载。
如有侵权,请联系 cloudcommunity@tencent.com 删除。
评论
登录后参与评论
推荐阅读
目录

